23 February 2021
DNA milestone: one step closer to completing the tree of life for plants
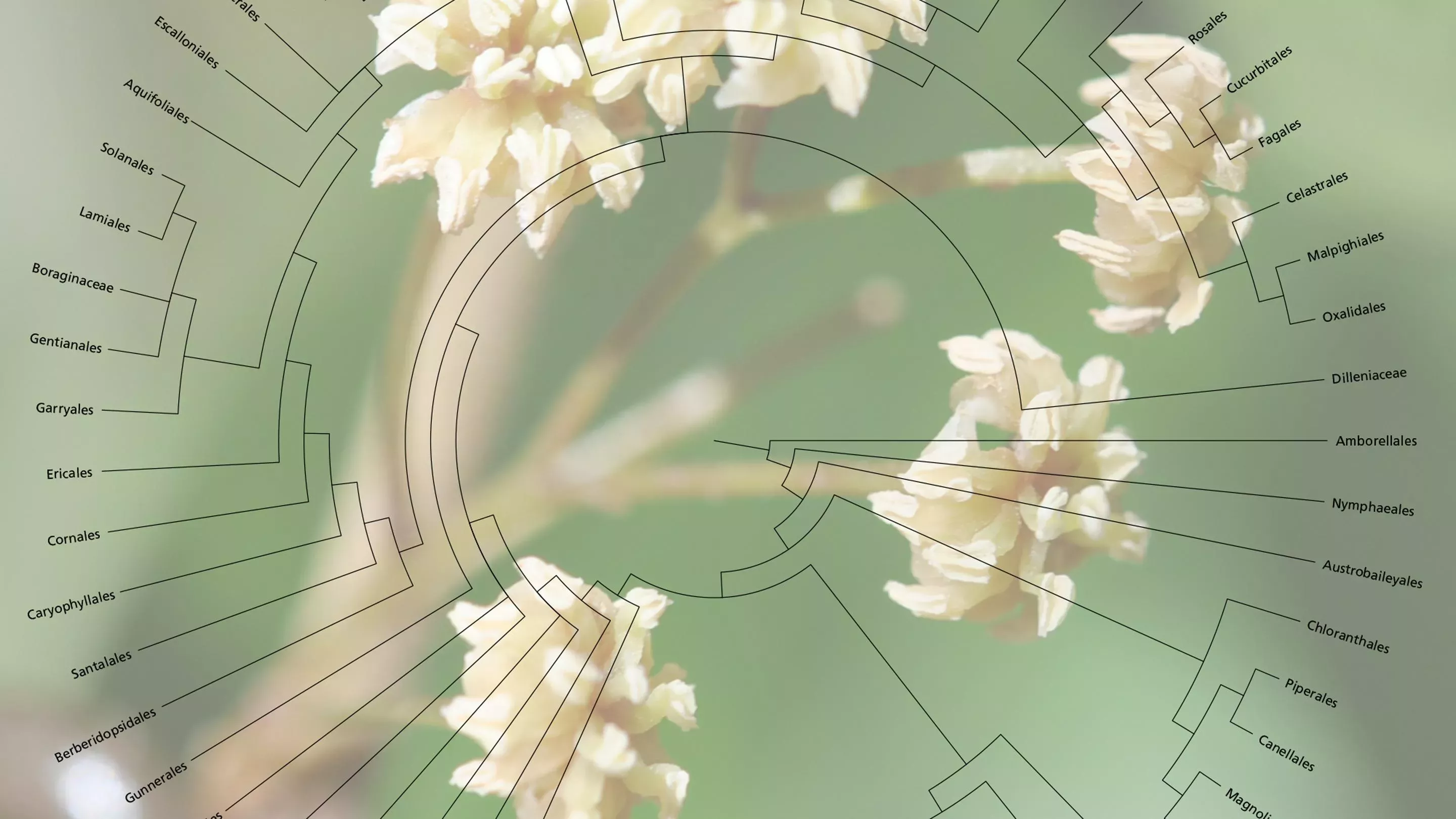
Understanding the evolutionary tree of life is the ultimate endeavour in biology, equivalent to constructing the periodic table in chemistry. Researchers at Kew are now well on the way to completing the tree of life for flowering plants by sequencing the DNA of one sixth of all genera.

Kew scientists have achieved a major milestone in a vast project to complete the trees of life for plants and fungi.
These trees of life, just like our own family tree, enable us to understand how different species are related to each other, and are vital for helping future species discovery, monitoring and conservation.
Botanical transformation
All life on Earth is related but working out how requires a detailed comparison of the DNA sequences that are passed on from generation to generation like a molecular record of evolutionary history.
Until recently, analysing DNA sequences from large numbers of specimens was simply not possible.
Instead, the relationships between different plants were estimated by looking at their physical and chemical characteristics and using fossil records to establish a timeline of connections.
In the past 25 years, however, the application of DNA analysis to botany has transformed our understanding of how plants are related to each other on the many branches of the tree of life.

DNA discoveries
DNA has also proven to be an essential tool in the discovery of new plant species.
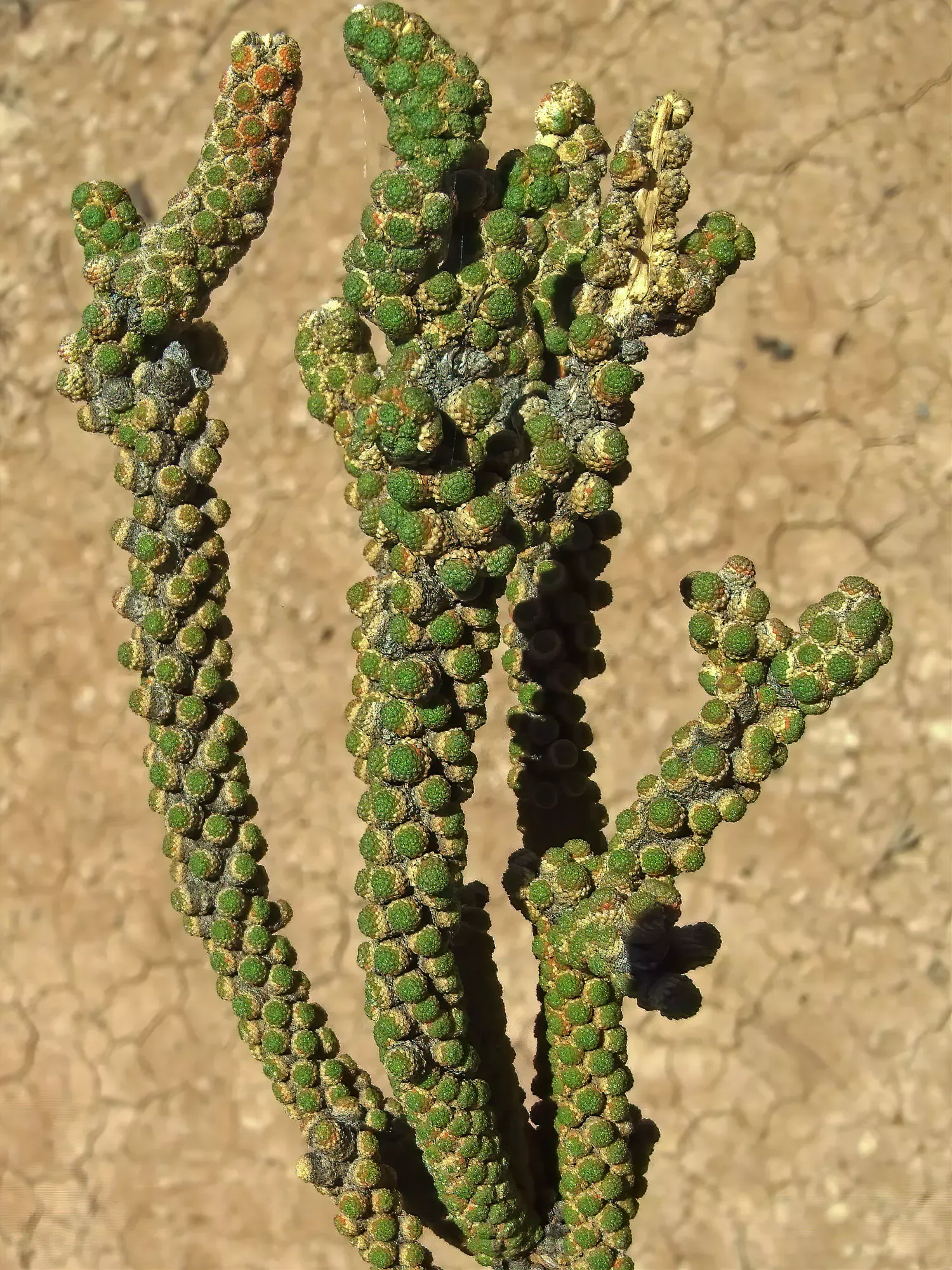
For example, back in 2010, a small inconspicuous, scaly shrub was discovered growing near salt pans in the Namibian desert.
It was so unlike anything else known to science that botanists were not able to place it confidently on the tree of life even in a family.
However, once DNA sequences were obtained, it was found to belong to a group of plants related to the cabbage and papaya families.
Its position on the tree of life was so unexpected, that it was assigned to its own new family, the Tiganophytaceae.
Without DNA evidence, it would have remained a mystery, creating an obstacle for future research or even conservation action.
Such revelations and discoveries inspired the creation of Kew’s newest garden, the Agius Evolution Garden, which lays out plants based on their evolutionary relationships discovered through their DNA.

Nevertheless, despite some technological advances and extensive collaborations across the botanical community, DNA information remained patchy for a long time, and by 2016 70% of species still lacked any DNA sequence data.
To draw a comprehensive map of the evolutionary connections across all species, the analysis of DNA needed to move up a gear – and this is where Kew’s flagship Plant and Fungal Trees of Life (PAFTOL) project came in.
In contrast to early DNA studies which analysed just one or a few genes to explore evolutionary relationships, PAFTOL uses hundreds of genes from each species in order to systematically build a comprehensive evolutionary tree.
This work is now possible thanks to a recent revolution in DNA sequencing technologies, enabling vast amounts of genetic information to be generated and deciphered at speed and ever decreasing cost.

Molecular "fishing"
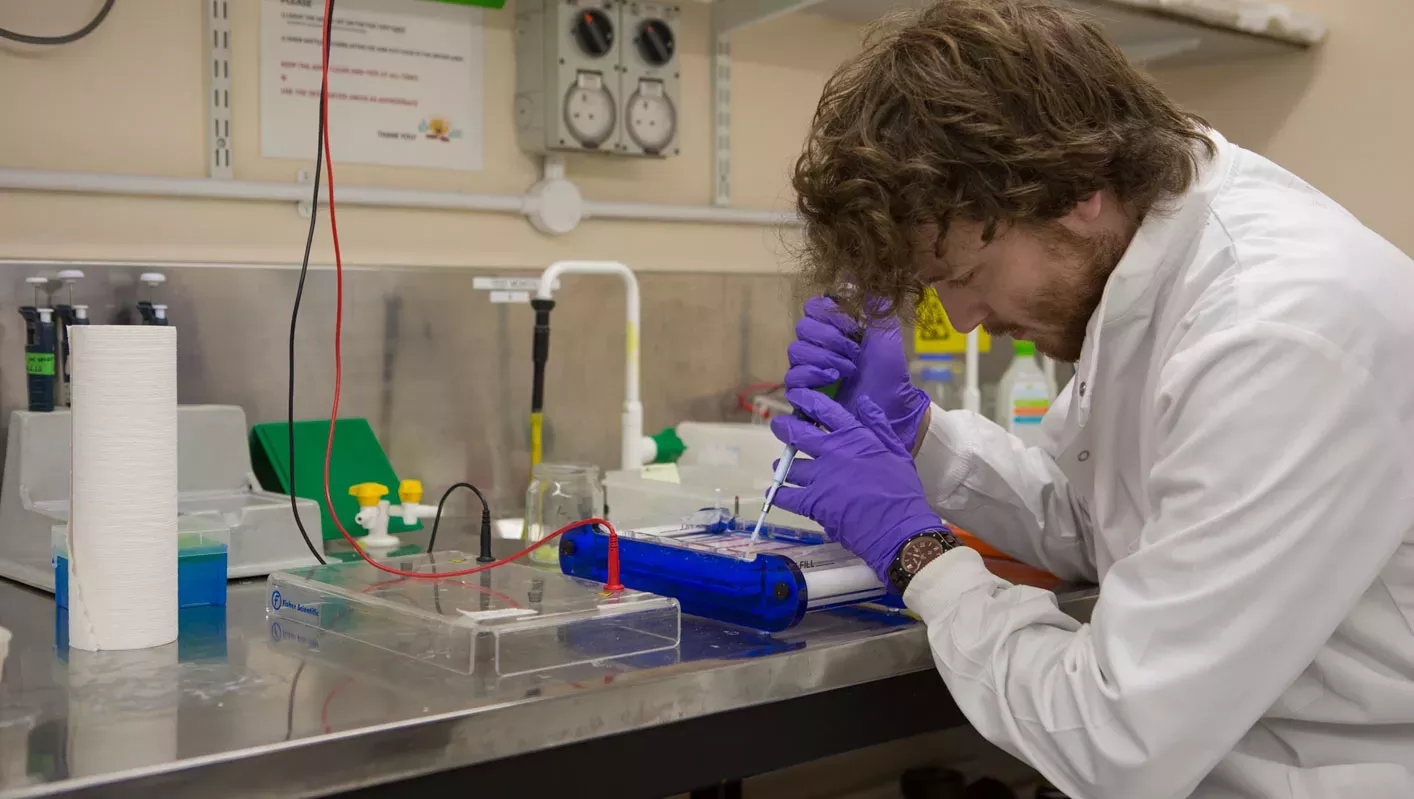
Kew has taken a lead in applying new sequencing advances at scale.
In collaboration with Chicago Botanic Garden, Texas Tech University, the One Thousand Plant Transcriptomes Initiative and others, its scientists have developed a novel toolkit, known as Angiosperms353, consisting of a set of molecular probes that can “fish out” 353 genes selected for their usefulness in building the tree of life for flowering plants (also known as angiosperms).
Using this toolkit, the Kew team has now gathered DNA sequence data for at least one species from 2,333 genera of flowering plants, representing 17% of the world’s 13,862 angiosperm genera.
It even includes 104 genera for which there was previously no sequence data at all.
This unprecedented dataset is being released online for scientists everywhere to study via a new web portal, the Kew Tree of Life Explorer.
From here, users can not only download and utilize the data for their own research but also explore evolutionary relationships using the tree of life generated from the new DNA sequence data.
Achieving this landmark required more than smart science.
Researchers also needed to access a huge breadth of biological samples, which Kew’s extensive collections helped provide.
In fact, around 80% of all samples analysed in the project were taken from Kew’s living collections, its DNA bank (a giant freezer in the Jodrell Laboratory set at minus 80 degrees C), the Millennium Seed Bank at Wakehurst, and specimens from Kew’s Herbarium, some of which were collected over 200 years ago.
Filling in the gaps
Completing this extensive programme of work in the middle of a pandemic has not been easy, with many scientists furloughed and key laboratories closed for lengthy periods.
However, having proved that the Angiosperms353 toolkit works on thousands of samples, the pace of analysis is expected to pick up from here and the Kew team has a target to sample 50% of all angiosperm genera by the end of 2021.
What is more, the fact that the DNA toolkit is openly available to the scientific community means that other institutions around the world can also conduct their own analysis.
For example, the Genomics for Australian Plants initiative has adopted Kew’s approach and is coordinating with PAFTOL to ensure the two teams cover maximum ground.
Where appropriate, such additional data will also be incorporated into the Kew portal to increase the sharing of this growing tree of life dataset as widely as possible.
As a result, a completed genus-level angiosperm tree of life is now well within reach. However, the work will not stop there.
The task of sampling all ~330,000 flowering plant species still remains to be done.
This is a 24-fold greater challenge than sampling the genera – but the good news is that the Angiosperms353 probe gives enough granular detail to identify variations between species and even, in some cases, within species.
Angiosperms are the most diverse group of land plants but, of course, completing the whole of the Plant and Fungal Trees of Life means scientists also need to sequence the non-flowering plants such as conifers and mosses – a challenge that is being tackled in parallel by U.S. institutions under the GoFlag project.

The unknown kingdom
And then there are the fungi, which may have smaller genomes than plants but still represent a daunting task.
Currently, there are estimated to be around 8,000 fungal genera, yet our understanding of their true diversity is still very limited compared with plants, and so this number may well grow considerably in the future.
While progress has not been as extensive as in plants, recent funding has enabled this daunting challenge to start, with a particular focus on sorting out the backbone of the fungal tree of life and exploring one of the most economically important fungal groups – the Ascomycetes.
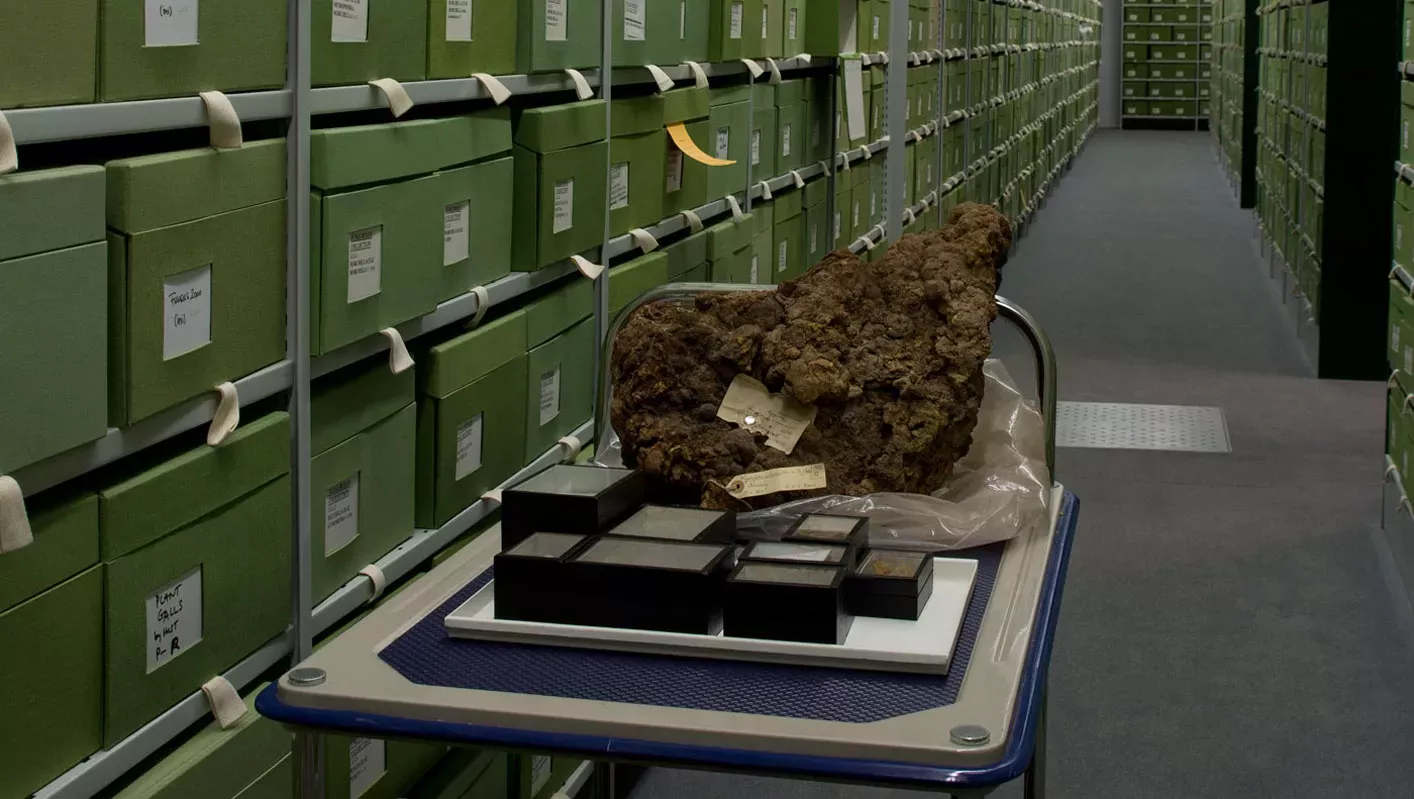
As for plants, this project will take advantage of the ability to extract DNA from the vast wealth of dried specimens in Kew’s Fungarium collections which are estimated to contain over 60% of fungal genera.
The new understanding of evolutionary relationships will be invaluable for guiding the exploration of fungal processes and products, with the potential to discover novel chemicals, antibiotics and biofuels.


Revealing secrets
One thing is clear: the pace of progress in unlocking the DNA secrets of the natural world is accelerating and it is now possible for science to address challenges that not so long ago would have seemed like science fiction.
These include the Earth Biogenome Project’s “moonshot for biology” plan to catalogue the genetic code of all known species of complex life within 10 years, and the Darwin Tree of Life Project which is already underway with the aim to sequence the complete genomes of all 70,000 British and Irish species.
All these projects are designed to provide a knowledge base to conserve, protect and restore biodiversity – and they would not be possible without the goldmine of resources stored in the world’s natural history collections, including Kew.
Read the paper
Baker W.J. et al. (2021) A Comprehensive Phylogenomic Platform for Exploring the Angiosperm Tree of Life. Systematic Biology, syab035
Acknowledgments
PAFTOL is currently supported by the Calleva Foundation and Evolution Education Trust and has previously received support from the Sackler Trust and the Garfield Weston Foundation.