4 January 2019
Completing the tree of life for plants and fungi
The tree of life allows us to discover, understand, utilise and conserve life on Earth. In this blog, Bill Baker explains how Kew is leading the charge in a new genomic age for tree of life research on plants and fungi.
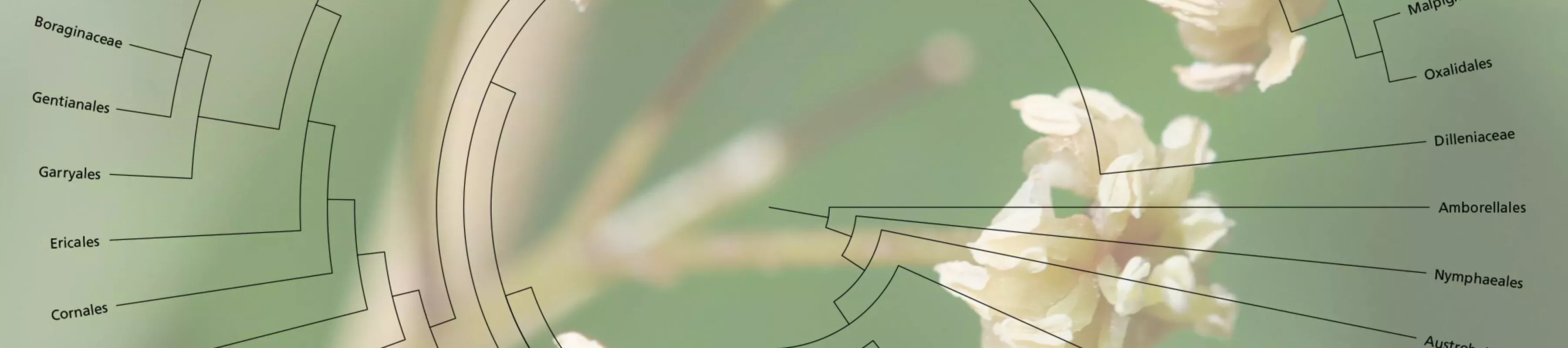

All living things are connected by their evolutionary history.
The evolutionary links between the Earth’s nine million species form a vast family tree - the tree of life.
The tree is like a roadmap to life on Earth - it helps us to discover, understand, utilise and conserve living things. Revealing this tree of life is one of the most fundamental challenges in science today.
Discovering the tree of life
We discover the tree of life by comparing DNA sequences between species.
Over the past two decades, Kew has taken a lead in this work, resulting in startling discoveries and a radical new classification of the flowering plants, which for the first time accurately reflects evolutionary history.
Nevertheless, genetic data have been collected from only about one quarter of the 391,000 known vascular plant species.
The challenge for fungi is even greater: millions of species remain to be discovered, let alone have their DNA analysed.

A new genomic age for tree of life research
A revolution in DNA sequencing technology is underway – whole genomes can now be deciphered at ever decreasing cost.
As a result, there is a growing movement globally to sequence the genomes of all species on Earth.
However, the main limiting step for such an ambition is not technology, but access to biological samples. For plants and fungi, this is where Kew comes in.
Kew holds the richest plant and fungal collections in the World. With these collections, we can make a globally important contribution to this movement in collaboration with our partners worldwide.


The Plant and Fungal Trees of Life Project at Kew
As a flagship project of our Science Strategy, Kew is completing the Plant and Fungal Trees of Life Project (PAFTOL).
To achieve this ambitious goal, we will generate and compile genome-scale data for at least one species of all 14,000 flowering plant genera and all 8,200 fungal genera.
PAFTOL Plants is well underway, with a 15-strong team of dedicated researchers.
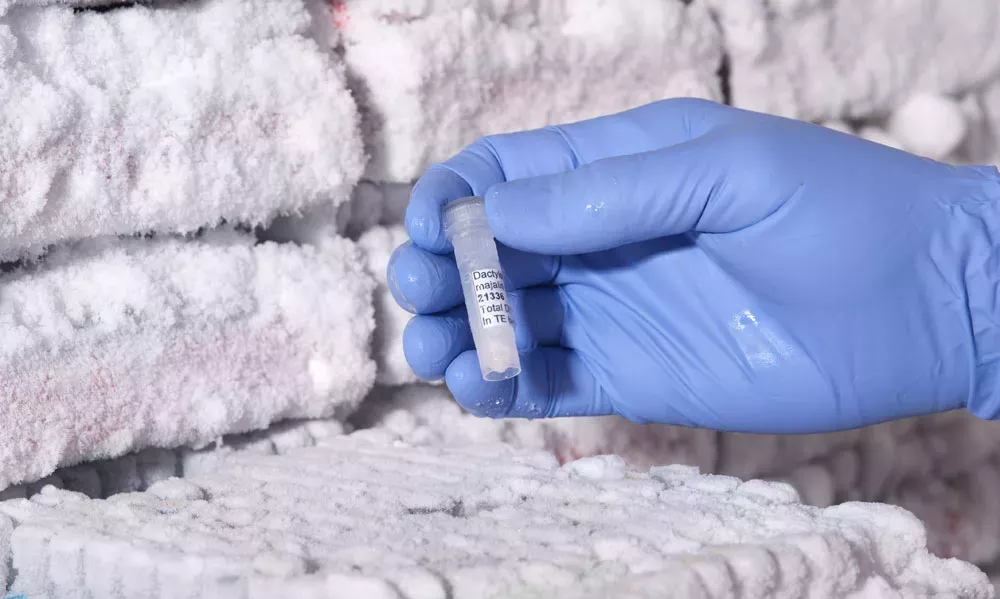
More than half of the samples we need are available in our living collections, seed bank and DNA bank. These are the collections that yield the best quality DNA for our labwork.
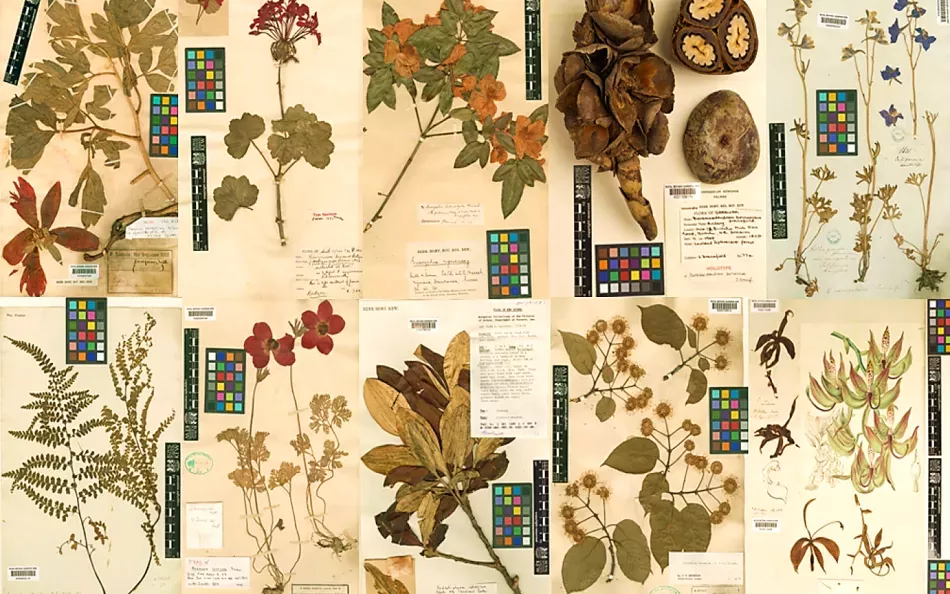
Gaps that exist in these ideal collections can be plugged thanks to Kew’s amazing herbarium collection. We now routinely sequence hundreds of genes from herbarium samples up to 200 years in age.
Our Victorian founders could not have imagined that their specimens would become an invaluable resource for deciphering the blueprint of life.
By the end of this year, we expect to have sequenced a full 25% of all flowering plant genera. To do this, we have created a novel genomic toolkit in collaboration with Chicago Botanic Garden, Texas Tech University and the 1000 Plants project.
The toolkit (known as Angiosperms-353) consists of a set of molecular probes that effectively “fish” out 353 genes selected for their usefulness in building the tree of life. This toolkit is now openly available to the scientific community and has the potential to develop as a community-wide standard for plant tree of life research.
The genomic era is dawning and Kew is embracing it. We are seizing this opportunity to unlock our unrivalled collections and expertise to break new ground in this urgent scientific endeavour.
References
Eiserhardt, W.L. et al. (2018). A roadmap for global synthesis of the plant tree of life. American Journal of Botany 105(3): 614-622.
Johnson, M., et al.(2018). A Universal Probe Set for Targetted Sequencing of 353 Nuclear Genes from Any Flowering Plant Designed Using k-medoids Clustering. Systematic Biology syy086.
Lewin, H.A., et al. (2018). Earth BioGenome Project: Sequencing life for the future of life. Proceedings of the National Academy of Sciences of the United States of America 115(17): 4325-4333.